Background
Neuroblastoma is the most common extracranial solid pediatric cancer accounting for 8-10% of cancers in childhood and 15% of pediatric oncology deaths. Neuroblastoma arises from the developing sympatho-adrenal lineage during the embryonic development. It is a genetically heterogeneous disease with a diverse clinical outcome ranging from spontaneous tumour regression to malignant metastatic disease with relapses and poor response to current therapy. While patients whose tumours undergo spontaneous regression or maturation (ganglioneuroblastomas, ganglioneuromas) have mostly an excellent outcome, only a minority of children with aggressive tumours can be cured. Despite the advances in genomic and trancriptomic analyses, the identification of molecular determinants of the very poor therapeutic response and worst outcome of high-risk patients remains challenging. Thus, a better understanding of the biology of both, spontaneously regressing/maturing and aggressive tumours is of high interest to develop novel treatment approaches.
Our research
Biology of high-risk neuroblastoma
One of our main research interests is the biology of high-risk neuroblastoma. Patients that are diagnosed and stratified as high-risk suffer from relapses and metastases and their survival rate remains below 40% despite intensive multimodal treatment. To date there are only a few driver genes linked to the pathogenesis of high-risk neuroblastoma, most of which are not directly druggable and frequently insufficient response to therapy is observed. In our group, we employ state-of-the-art technologies, such as genome-wide and targeted CRISPR/Cas9 screens and single cell genomics and epigenomics in order to identify the oncogenic drivers and epigenetic dependencies in tumours from high-risk neuroblastoma patients. We have established in vitro and in vivo preclinical patient-derived models for functional assays and drug testing for precision oncology that can be translated into existing and new clinical trials with the ultimate goal to improve treatment outcomes and survival of high-risk neuroblastoma patients.
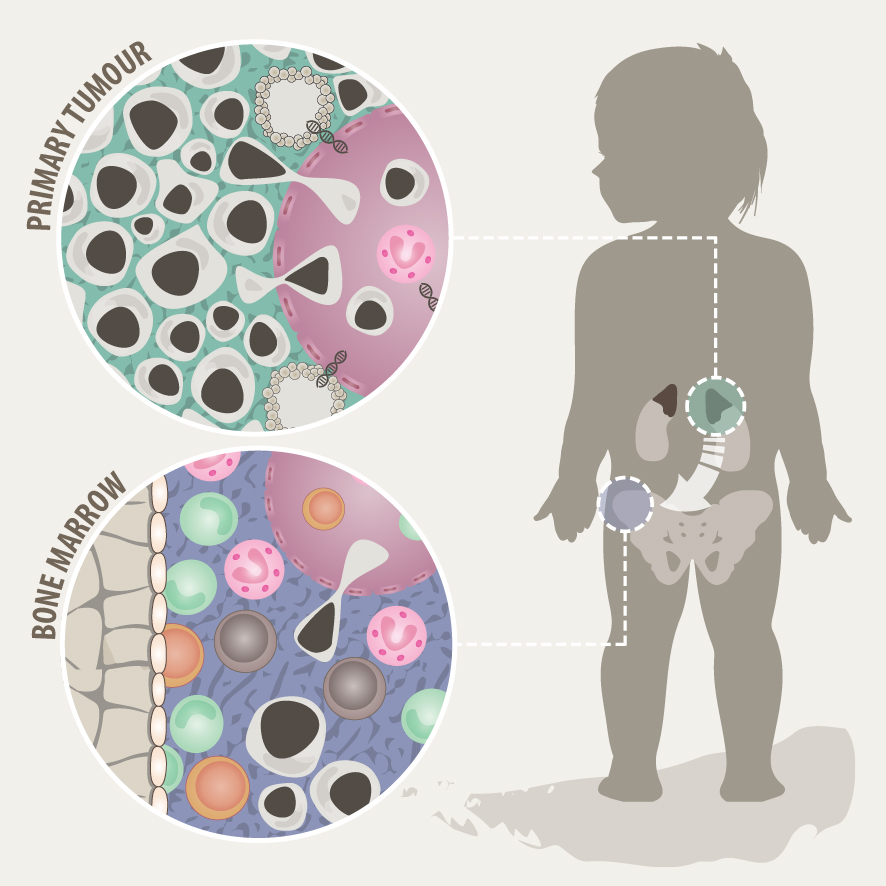
Tumour heterogeneity and microenvironment
Solid tumours often consist of different subpopulations of cells that harbor distinct genotypes and phenotypes. This results in a variation of clinically important features such as the abundance of prognostic markers and therapeutic targets, leading to differential levels of treatment sensitivity. Tumour cell metastasis and adaptation to new tissue microenvironments can further promote inter- and intratumour heterogeneity among metastasizing and disseminated tumour cells. In support of this notion, we have recently shown that disseminated tumour cells in the bone marrow substantially differ from the tumour they originated from in regards to their genetic makeup and expression programs. Tumour cells disseminate to the bone marrow in various solid cancers such as neuroblastoma, breast cancer and Ewing sarcoma, which is associated with poor outcome. In the majority of metastatic neuroblastoma patients, disseminated tumour cells are present in the bone marrow already at the time point of diagnosis. Our aim is to capture the full spectrum of tumor cells in neuroblastoma and to understand their interaction with the tumor microenvironment at the primary site and in the metastatic bone marrow by using novel single-cell-omics and multiplex imaging technologies. This will allow us to identify new biomarkers and to develop better therapeutics for targeted treatment.
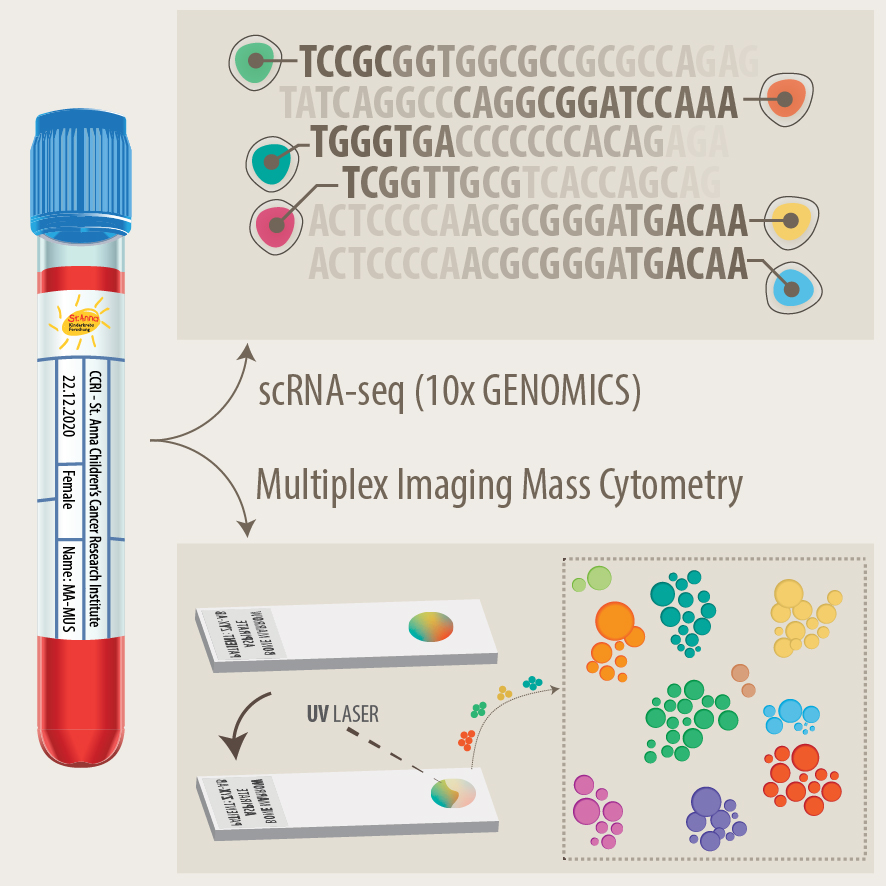
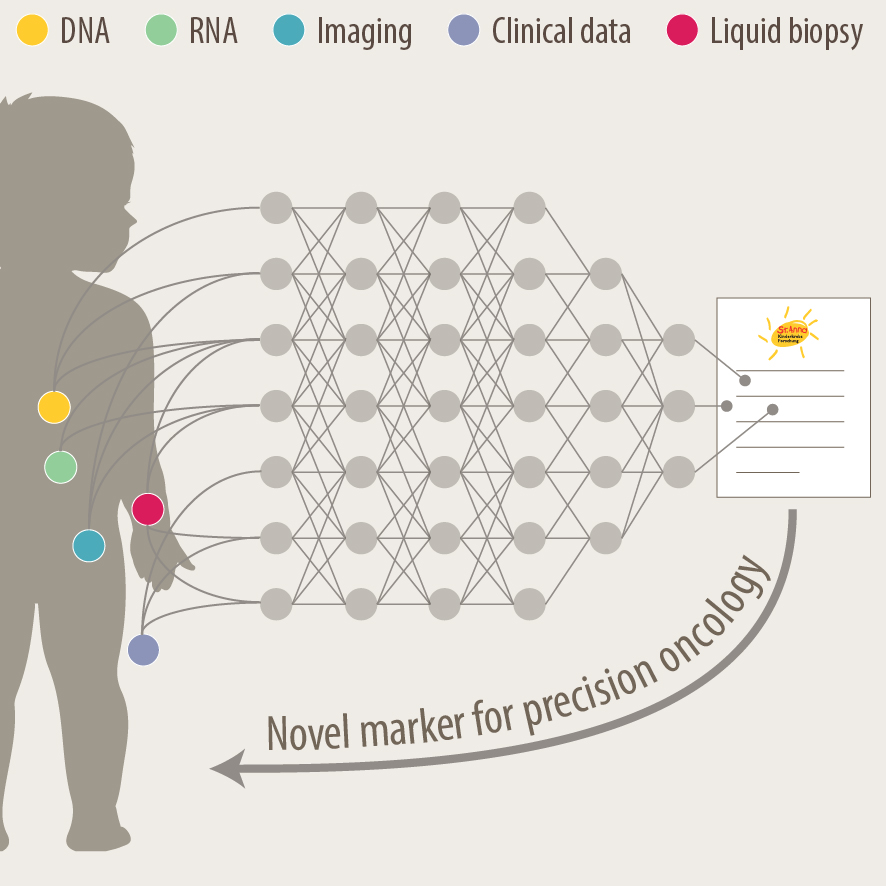
Development of new diagnostics and prognostic markers for precision oncology
Another focus of our group is the translation of current research to clinical practice with the development of better diagnostic approaches and prognostic markers. As pediatric solid tumours are rare, this can only be addressed within the scope of multi-center as well as multi-disciplinary cooperation. Towards this, we are part of different consortia and collaborative studies, that bring together experts in the fields of biological and computer-based research with pediatric oncologists. In addition to molecular profiling of the primary tumor and bone marrow, novel liquid biopsy approaches, i.e. the analysis of tumor markers in body fluids, are important tools to monitor cancer patients and detect relapse early. We employ advanced bioinformatics analyses, AI-based machine-learning and customized visualization strategies on complex multi-dimensional data for identifying novel markers for precision oncology. As an example, we have recently developed the VISIOMICS software platform http://www.visiomics.at/, which supports an integrated analysis of image and multiOMICS data for tumour diagnostics.
- We work in close collaboration with the Tumour Biology Diagnostic Team, which offers diagnostic services as ISO9001 certified and national reference lab for several ongoing clinical studies. For more information please see: Tumor Biology – Labdia.
- CCRI Biobank for pediatric solid tumours in collaboration with Labdia.
- Automated imaging devices
Projects and Funding

- A SIOPEN pragmatic clinical trial to MOnitor NeuroblastomA relapse with LIquid biopsy Sensitive Analysis (MONALISA)
CCRI responsible Principal Investigator and Scientific Coordinator: Sabine Taschner-Mandl
Coordinator: SIOPE, Belgium
Grant from the European Commission, Horizon Europe, ID – 101137028
Duration: 01/01/2024 – 31/12/2028

- MAPMET – Mapping metastatic cancer by multi-modal imaging
CCRI responsible Principal Investigator: Sabine Taschner-Mandl
Grant from the Austrian Science Fund (FWF), Stand-Alone Project, DOI: 10.55776/P35841
Duration: 01/05/2022 – 30/04/2026
- EXPLORE-NB
CCRI responsible Principal Investigator: Sabine Taschner-Mandl
Grant from the Austrian Science Fund (FWF), International – Multilateral Initiative: ERA-NET/Transcan-3, DOI: 10.55776/PIN2827223
Duration: 01/11/2024 – 30/10/2027
- DART2OS – Devising Advanced TCR-T cells to eradicate OsteoSarcoma
CCRI responsible Principal Investigator: Sabine Taschner-Mandl
Grant from the Austrian Science Fund (FWF), Emerging Fields, DOI: 10.55776/EFP45
Duration: 01/10/2024 – 30/10/2029

- Prognostic impact of genome wide copy number and tumor cell state on patients enrolled in the SIOPEN/HR-NBL1 tria
CCRI responsible Principal Investigator: Sabine Taschner-Mandl
Grant from SIOPEN (Grant Call 2024)
Duration: 01/06/2025 – 31/05/2027
Selected Articles
Collaborating Partners
National: Nikolaus Fortelny (University Salzburg, Austria), Christoph Bock (CeMM/BSF, Austria), Katja Bühler (VRVis, Austria), Alan Hanbury (Technical University Vienna, Research Studios Austria), Lukas Fischer (Software Competence Center Hagenberg, Austria), Michael Brandstötter (CogVis, Austria), Christopher Gerner (University of Vienna, Austria) International: Hedwig Deubzer (Charite, Germany), Gudrun Schleiermacher (Institute Curie, France), Lieve Tytgat (Princess Maxima Center, Netherlands), Jan Molenaar (Princess Maxima Center, Netherlands), Frank Westermann (DKFZ, Germany), Jaime Font deMora (Research Institute Hospital La Fe, Spain), Adela Canete (University Hospital La Fe, Spain), Barbara Hero (University Clinic Cologne, Germany), Christian Ostalecki (University Clinic Erlangen, Germany), Jelena Racson (Vilnius University Children’s Hospital, Lithunia), Bernd Bodenmiller (ETH Zurich and University of Zurich, Switzerland) Collaborating clinicians at St. Anna Children’s Hospital and Vienna General Hospital: Ruth Ladenstein, Stefan Fiedler, Caroline Hutter, Elisabeth Salzer, Heidrun Boztug, Stefan Köhrer, Leo Kager, Martin Metzelder
About Sabine Taschner-Mandl
Sabine Taschner-Mandl, PhD has been head of the Tumor Biology group at St. Anna Children’s Cancer Research Institute, CCRI, since 2018, where she has been working as a scientist since 2008. The aim of her research group is to tackle unresolved questions of neuroblastoma pathogenesis and develop new diagnostic and therapeutic approaches to facilitate precision medicine for children with malignant tumors. In addition, Sabine Taschner-Mandl is a lecturer at the Medical University of Vienna and Vienna University of Technology. She completed her studies of biology at the University of Vienna with a diploma thesis in vaccine development. This was followed by a dissertation and a postdoc position at the Institute of Immunology at the Medical University of Vienna. Besides her research work at CCRI, the researcher was a visiting scientist at Significo and the University of Helsinki (EC-FP7 Marie Curie Program). She is leading and participating in several international and national projects and initiatives. As member of the Executive Board and co-chair of the Biology Committee of the European Society of Pediatric Oncology Neuroblastoma (SIOPEN) and in other international working groups (INRG, ITCC), she is fostering innovative research for the benefit of pediatric patients with cancer.













